ABCdeZ.jl
Approximate Bayesian Computation (ABC) with differential evolution (de) moves and model evidence (Z) estimates.
ABCdeZ.jl offers Bayesian parameter estimation and model comparison/selection for inference problems with an intractable likelihood. Models only need to be simulated (instead of calculating the likelihood). In this documentation you will find everything to get started.
ABCdeZ.jl was developed @TSB by Maurice Langhinrichs and Nils Becker. This work is based on many people's previous achievements, particular some part of the code base was adapted from KissABC.jl [1]; please find a complete list of references below.
Brief introduction
Bayesian inference allows to update a parameter prior $p(θ | M)$ in the light of new data $D$ to obtain posterior knowledge as
\[p(θ | D, M) = \frac{p(D| θ, M) \, p(θ | M)}{Z}\]
where $θ$ are the parameters of model $M$, $p(D| θ, M)$ the likelihood and $Z = p(D | M)$ is the model evidence (also known as marginal likelihood). The model evidence is the normalising integral
\[Z \equiv p(D | M) = \int \, p(D| θ, M) \, p(θ | M) \, \mathrm{d}θ.\]
Next to parameter estimation, Bayes' theorem also applies to the outer layer for model selection/comparison, updating (by the same data) model prior probabilities $p(M)$ to posterior probabilities as
\[p(M | D) = \frac{p(D | M) \, p(M)}{p(D)}\]
where the model evidences reappear as the essential factors, and $p(D) = \sum_i p(D | M_i) \, p(M_i)$ is a normalisation factor over all models $M_i$ ($i = 1,...,n$).
ABC (Approximate Bayesian Computation) approximates the likelihood through model simulations [2], yielding simulated data $D'$ that are evaluated with a kernel $π_ϵ(D' | D)$,
\[\hat{p}(D| θ, M) = \int \, p(D' | θ, M) \, π_ϵ(D' | D) \, \mathrm{d}D',\]
where $ϵ ≥ 0$ specifies the width of the kernel. Under certain conditions, it can be shown that this approximation converges for $ϵ \rightarrow 0$ [2], keeping only model simulations $D'$ that are "very close" to the original data $D$.
These equations show in brief, that posterior parameter and posterior model probabilities can be obtained by updating (model and parameter) priors through some data $D$, only requiring model simulations and a sufficiently small target $ϵ\approx 0$.
ABCdeZ.jl will provide parameter samples approximating the posterior $p(θ | D, M)$ and estimations of (logarithmic) model evidences $\mathrm{log}Z$.
ABCdeZ.jl
ABCdeZ.jl currently implements two main ABC methods:
abcdesmc! (from "abc de smc"): A Sequential Monte Carlo algorithm. For posterior samples and model evidence estimates. Method of choice for multimodal or complex posterior landscapes.
abcdemc! (from "abc de mc"): A simpler MCMC algorithm. For posterior samples only. "Greedy" algorithm that may be faster than
abcdesmc!for some problems. May have difficulties with multimodal problems (useabcdesmc!instead).
Both methods make differential evolution (de) proposals for the MCMC (Markov chain Monte Carlo) steps of the algorithms; differential evolution provides a good scale and orientation to explore the (often) complex multi-dimensional parameter spaces effectively [4]. For more information on both algorithms follow the respective links above.
Why ABCdeZ.jl?
- If you want to perform parameter estimation and/or model comparison, but it is difficult to compute the likelihood for your models (time-consuming and/or analytically unknown). ABC is a likelihood-free approach, the only requirement is that you can simulate your models, given values for their parameters.
- ABCdeZ.jl allows to compute model evidences directly and hence the results are future-proof. Model selection in ABC is often done by simulating all models of interest in the same ABC run. With ABCdeZ.jl you can run models individually, store the resulting evidence (and $ϵ$'s) and compute evidences of any other models later, without re-computing the first. In the end, the desired set of models can be compared by transforming their evidences into posterior model probabilities or Bayes factors.
- ABCdeZ.jl offers fast thread-based parallelism enabled by FLoops.jl [10]. Additionally it also allows to store arbitrary data
blobstogether with the sample particles (e.g., to have the simulation output alongside the final posterior samples). For more information see here.
Why not ABC(deZ.jl)?
- If you can calculate the likelihood function for your models, make use of it. Instead of ABC, consider likelihood-based inference methods.
- Please be aware of the caveats of ABC, particular for model comparison/selection. ABC is approximative and may lead to wrong/misleading results if not applied carefully (see here for more).
Minimal example
Here we will use ABCdeZ.jl to infer the model evidences and posterior distributions within a simple toy example. The models have a single parameter $θ$ that is, a priori, normally distributed (prior). After seeing one single data point, we obtain an updated posterior knowledge. To obtain model evidences, next to the posterior samples, we will use the abcdesmc! method; for posterior samples only one may also use abcdemc! with a very similar syntax (as seen here).
ABCdeZ.jl (as ABC in general) requires model simulations (for samples of $θ$) only, and not the likelihood values (at these $θ$). However, in this simple example the likelihood is available and chosen in a way (also Normal) that the prior is conjugate, and the exact posterior distributions and evidences are known analytically. We will compare the inferences made by ABCdeZ.jl with the analytical counterparts.
We load the required packages, set the single data point and the target $ϵ$ for the ABC runs.
using ABCdeZ
using Distributions
### data
data = 3
### ABC target ϵ (maximum distance)
ϵ = 0.3Then we set up the inference of a first model. The normal prior is specified via Distributions (for more see "Prior" box below). Note that the model is solely specified by a random simulation for a given $θ$. The distance function here simply reports the absolute distance between the random model output and the single data point (see here for additional features in the distance function).
### model 1 inference
σ₁² = 10
prior1 = Normal(0, sqrt(σ₁²))
# model simulation (to replace likelihood)
model1(θ) = rand(Normal(θ, 1))
# distance function between model and data
dist1!(θ, ve) = abs(model1(θ)-data), nothingWith the following line we run the abcdesmc! method of ABCdeZ.jl. The inference result is stored in r1.
### ABC run
# run the smc method for model 1
r1 = abcdesmc!(prior1, dist1!, ϵ, nothing,
nparticles=1000, parallel=true)From the result r1 we can read out the posterior samples (important: weighted by Wns, see box "Posterior sample weights" here) and the estimated model evidence for model 1 (transforming back from log-scale).
### process results
# posterior parameters
posterior1 = [t[1] for t in r1.P[r1.Wns .> 0.0]]
# model evidence (logZ is the logarithmic evidence)
evidence1 = exp(r1.logZ)Plotting the posterior1 samples as a histogram, one can see the gained knowledge from the prior just by the single data point (Figure below, left panel). The samples also match the analytical posterior, available in this simple example.

To demonstrate model comparison enabled by ABCdeZ.jl, we now repeat the procedure with a second model. Here, for simplicity, model 2 only differs from model 1 in the prior; of course in more interesting settings, not only the prior, but the whole architecture of models may be different (kind of and/or number of parameters). Any models can be compared in principle; as long as inferences are done for the same (summary) data (and simulations always match the structure of the data), distance method (here abs()) and target $ϵ$. The same target $ϵ$ is important in ABCdeZ.jl due to the (typically) unnormalised ABC kernel (see box "Model evidence (off by a factor)" here; if not possible, advanced info here).
### model 2 inference
σ₂² = 100
prior2 = Normal(0, sqrt(σ₂²))
model2(θ) = model1(θ)
dist2!(θ, ve) = abs(model2(θ)-data), nothing
r2 = abcdesmc!(prior2, dist2!, ϵ, nothing,
nparticles=1000, parallel=true)
posterior2 = [t[1] for t in r2.P[r2.Wns .> 0.0]]
evidence2 = exp(r2.logZ)The posterior inference of model 2 is visually very similar to model 1, except the difference in the prior (Figure above, right panel).
Finally, the estimated evidences can be used to compute posterior model probabilities (based on equations here). The model prior is uniform between the two models here.
### model probabilities
# model priors (uniform here)
mprior1 = 0.5
mprior2 = 0.5
# model posterior probabilities
mposterior1 = evidence1*mprior1 / (evidence1*mprior1 + evidence2*mprior2) # posterior prob. model 1
mposterior2 = evidence2*mprior2 / (evidence1*mprior1 + evidence2*mprior2) # posterior prob. model 2The model probabilities can be visually compared (Figure below), recovering the exact analytical results. Note that evidence values by ABCdeZ.jl are numerically uncertain (see box "Uncertainty of model evidence" here); as such the figure below also shows evidence values from repeated runs.
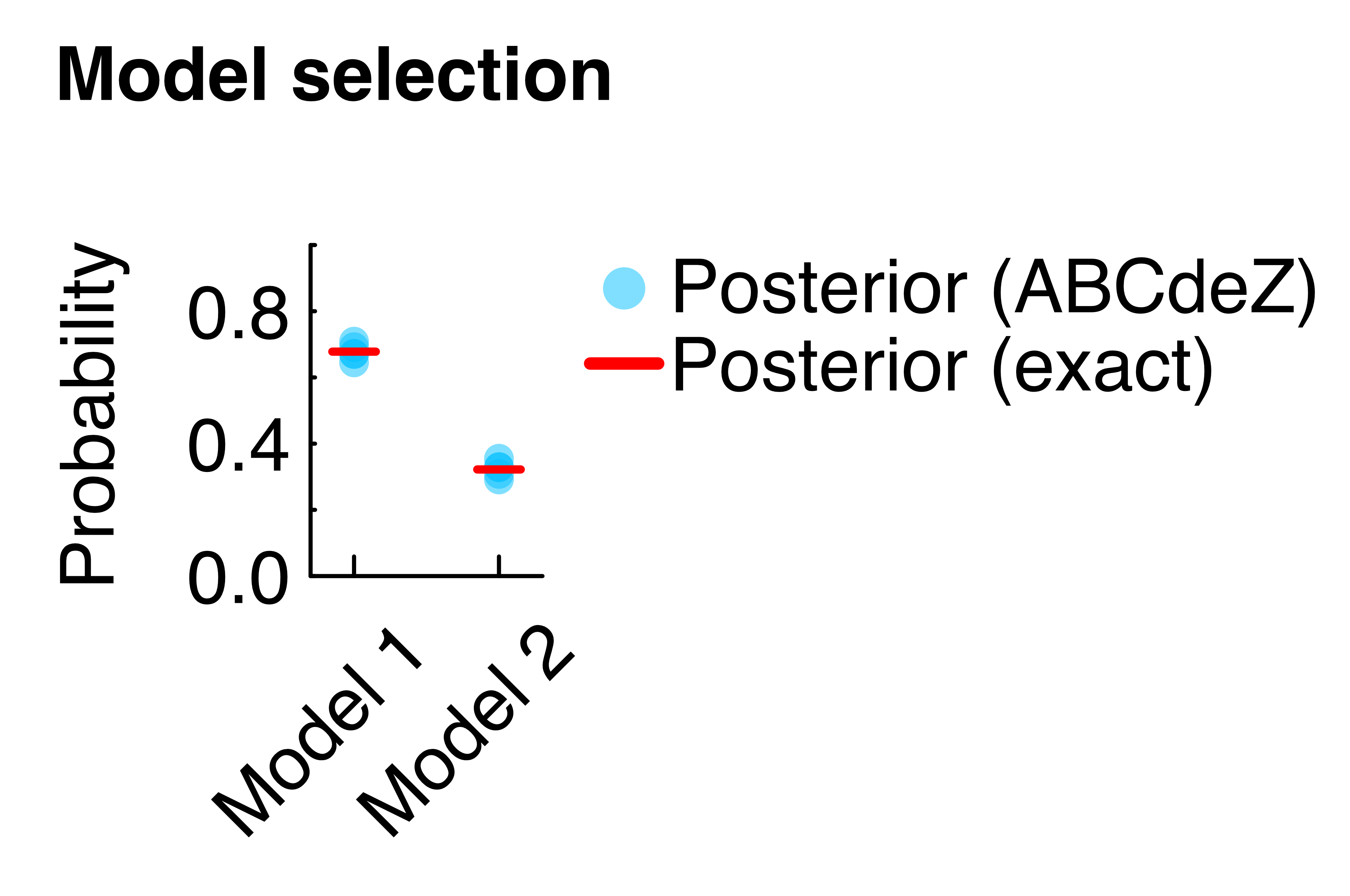
The complete code for this minimal example, including the derivation of the analytical counterparts, can be found on GitHub.
Multidimensional prior distributions (continuous, discrete or mixed) can be specified via the Factored() syntax (from independent 1d marginals), e.g. prior2d = Factored(Normal(0, sqrt(10)), DiscreteUniform(1, 10)).
Inference by abcdesmc!
Missing docstring for abcdesmc!. Check Documenter's build log for details.
Posterior samples obtained by abcdesmc! have to be associated with their weights (r.Wns). With an indicator ABC kernel (default) there are just two weights (i.e. alive and dead particles) and the correct posterior samples are hence given by posterior = [t[1] for t in r.P[r.Wns .> 0.0]] (for the first parameter here).
The model evidence estimates from the abcdesmc! method obtained by the default ABC indicator kernel are off by a normalisation factor coming from an unnormalised kernel (the one used in the final iteration). To do model selection/comparison this means that evidence estimates for the set of models have to be done for the same data (or summary statistics), distance function, ABC kernel and the same target ϵ (which is ϵ_target if run not stopped by nsims_max). Then the (unknown) normalisation factor is the same for all models and does not matter (cancels) for Bayes factors or posterior model probabilities. See here for workarounds if ϵ is not the same.
As of now the abcdesmc! method does not provide a (numerical) uncertainty for the model evidence estimate from a single run. It may be however very useful to check for this when doing model comparison (as the resulting Bayes factors or posterior model probabilities are uncertain as well). So, if the runtime permits, run the abcdesmc! method multiple times and collect the resulting set of evidence values for mean/median and std estimates.
Inference by abcdemc!
Missing docstring for abcdemc!. Check Documenter's build log for details.
The abcdemc! method implements a "greedy"/biased Metropolis-Hasting step in the Markov chain. This allows a fast convergence, particularly well-suited for unimodal problems. However, to obtain valid posterior estimates the algorithm needs to converge (all particles below ϵ_target and r.reached_ϵ==true). Otherwise the samples will be biased (closer to the MAP (maximum a posteriori probability) parameter values with reduced variation).
In contrast to the abcdesmc! method the resulting posterior samples of the abcdemc! method are not associated with weights and can be used directly, i.e. posterior = [t[1] for t in r.P] (for the first parameter here).
Distributions and Priors
Missing docstring for Factored. Check Documenter's build log for details.
Missing docstring for pdf. Check Documenter's build log for details.
Missing docstring for logpdf. Check Documenter's build log for details.
Base.rand — Functionrand(rng::AbstractRNG, factoreddist::Factored)Function to sample one element from a Factored object.
Base.length — Functionlength(p::Factored)Returns the number of distributions contained in p.
Various notes
ABC - Approximations
ABC is an approximate method, with approximation errors coming from typically two sources [2]:
The kernel width $ϵ$ (or target distance of an indicator kernel). Except for some discrete problems (where exactness may be reached with ABC and $ϵ=0$), $ϵ$ is chosen $>0$ which introduces an approximation error (see intro). Hence, the choice of $ϵ$ is a trade-off between accuracy and runtime.
Summary statistics instead of the complete data. Often data and model simulations are both reduced to some summary statistics (e.g., mean values) before they are compared with a distance function. If the summary statistics are sufficient (for parameter estimation or, more stringent, model comparison), $ϵ$ remains the only approximation. However, often the summary statistics are not sufficient, causing another level of inaccuracy that needs to be checked/controlled for. This is particularly important for model comparison (also see below and [2] [7] [8] and many more in the literature).
More on model evidences
Effect of summary statistics on model selection: Summary statistics may introduce a second source of approximation error. This needs to specifically kept in mind for model selection, as results may become entirely uninformative or wrong [2] [7]. Note that a summary statistic may be sufficient for parameter estimation for two models individually, but can be entirely useless for model selection between these two models [2]. It is hence critical to choose summary statistics that are sufficient or at least near-sufficient for the task of model selection. Marin et al. (2014) [7] provide a procedure to control for this (also more briefly described on stackexchange [8]).
Comparison of models with different final ϵ: Models should be compared based on evidence values computed for the same final $ϵ$ (due to kernel normalisation factor, as mentioned here or in the minimal example). Still, there may be cases where this is not possible. There are two potential workarounds:
Run the
abcdesmc!method with optionverboseout=true. Then the inference resultrwill contain the complete list of (logarithmic) evidence values (r.logZs) for each sequential $ϵ$ (r.ϵs). For models with different final $ϵ$ (r.ϵ), but $ϵ$ values in the list that are somewhat similar, one may use the smallest similar $ϵ$ and the respective evidence values for each model to compare them.For a model $B$ that seems worse than a current best model $A$, it may be hard to run ABC for model $B$ targeting the same small $ϵ_A$ of model $A$. A conservative evidence value for model $B$ is the evidence obtained for another higher $ϵ_B > ϵ_A$ (artificially favoring $B$ compared to $A$ at $ϵ_A$). Using such evidences for the (qualitative) task of model selection is fine, as long as model $B$ is still defeated by model $A$, as one does not make misleading conclusions.
Features for the distance methods
In the distance function in the minimal example (
dist!(θ, ve) = abs(model(θ)-data), nothing)veare "external variables" (varexternalinabcdemc!andabcdesmc!) that can be used in the distance computation and mutated in-place, even in the parallel mode (each thread will obtain its own copy for thread-safe parallel ABC runs).veis passed as 4th positional argument toabcdesmc!(nothingin the minimal example).In the distance function in the minimal example (
dist!(θ, ve) = abs(model(θ)-data), nothing) the second return argument (nothing) can be used to store arbitrary data (blobs) to each particle; theseblobswill be associated with the final posterior samples/particles in the end. For exampleblobscould record the actual simulation output:function dist2!(θ, ve, constants, data) # constants can be used to pass thread-safe constants that are NOT mutated; # ve for in-place, mutatable variables # distance method simdata = model(θ) blob = simdata d = abs(simdata-data) d, blob end dist2!(θ, ve) = dist2!(θ, ve, nothing, data) r = abcdesmc!(prior, dist2!, ϵ, nothing, nparticles=1000, parallel=true) posterior = [t[1] for t in r.P[r.Wns .> 0.0]] evidence = exp(r.logZ) blobs = r.blobs[r.Wns .> 0.0]
References
- Some part of the code was copied, adapted and/or inspired by KissABC.jl [1]. For example, the
Factoredsyntax was adopted,abcdemc!is based onABCDE,abcdesmc!is loosely based onsmc. We thank the developers of the package. - A very good theory background for the general approach of model evidences from single ABC runs is given by Didelot et al. (2011) [2]. More details on algorithms (in the likelihood-context) is found in Llorente et al. (2020) [3].
- The differential evolution moves are introduced in Ter Braak (2006) [4].
- As done also in KissABC.jl [1], the implementations of the
abcdemc!method are a simplified version of the method in Turner et al. (2012) [5]. The algorithmic idea inabcdesmc!is mostly based on Amaya et al. (2021) [6], next to KissABC.jl, particular the handling of weights and the adaptive differential evolution move tuning (Amaya et al. (2021) is in the likelihood context, which we adapted to ABC). - Closer read on sufficient summary statistics for model comparison in ABC is found in Marin et al. (2014) [7] and condensed in this stackexchange post [8].
- Stratified resampling (for
abcdesmc!) is inspired by Douc et al. (2005) [9]. - For fast thread-based parallelism we make use of FLoops.jl [10].
Index
- 1KissABC (https://github.com/francescoalemanno/KissABC.jl)
- 2Didelot et al. "Likelihood-free estimation of model evidence." Bayesian Anal. 6 (1) 49 - 76, 2011.
- 3Llorente et al. "Marginal likelihood computation for model selection and hypothesis testing: an extensive review" arXiv:2005.08334 [stat.CO], 2020.
- 4Ter Braak. "A Markov Chain Monte Carlo version of the genetic algorithm Differential Evolution: easy Bayesian computing for real parameter spaces" Statistics and Computing volume 16, pages 239–249, 2006.
- 5Turner et al. "Approximate Bayesian computation with differential evolution" Journal of Mathematical Psychology, 2012.
- 6Amaya et al. "Adaptive sequential Monte Carlo for posterior inference and model selection among complex geological priors" arXiv:2105.02012 [physics.geo-ph], 2021.
- 7Marin et al. "Relevant statistics for Bayesian model choice" J. R. Statist. Soc. B, 2014.
- 8https://stats.stackexchange.com/questions/26980/abc-model-selection
- 9Douc et al. "Comparison of Resampling Schemes for Particle Filtering" arXiv:cs/0507025 [cs.CE], 2005.
- 10FLoops (https://github.com/JuliaFolds/FLoops.jl)