Bifurcation Analysis
A numerical study of the changes in the dynamics and stability of a system upon variations in its parameters.
Procedure for stability analysis at fixed points
Consider the following system of ordinary differential equations:
\[\dfrac{dx}{dt} = F(x)\]
Determine the fixed point vector, $x^*$, solving $F(x^*) = 0$
Construct the Jacobian matrix, $J(x) = \dfrac{\partial F(x)}{\partial x}$
Compute eigenvalues of $J(x^*)$: $|J(x^*) − λE| = 0$
Conclude on stability or instability of $x^*$ based on the real parts of eigenvalues
- All eigenvalues have real parts less than zero → $x^*$ is stable
- At least one of the eigenvalues has a real part greater than zero → $x^*$ is unstable
Usage
Here I would like to use a mathematical model of Rb–E2F pathway (Yao et al., 2008) to show you how to perform bifurcation analysis with BioMASS.jl.
Prepare name2idx/ to define model species and parameters
- See examples.
Create set_model.jl
In this file, you will need to prepare four functions:
diffeq!: Ordinary differential equations of the model.param_values: Model parameters.get_derivatives: $\dfrac{\partial F(x)}{\partial bp}$, where $bp$ is the bifurcation parameter (x-axis).get_steady_state: Function to equilibrate the system.
Run create_diffeq function
create_diffeq(".")Then you will get forwarddiff.jl file in your model folder.
Load requirements
using DelimitedFiles
using Sundials
using SteadyStateDiffEq
using PyPlot
include("./name2idx/parameters.jl")
include("./name2idx/species.jl")
include("./set_model.jl")
include("./forwarddiff.jl")
const BP = C.S # name(index) of bifurcation parameter (x-axis)
const SN = V.NUM # num of state variables
const PN = 1 # num of parameters
const VN = SN + PN # num of variablesCalculate fixed points and analyze their stability
After executing new_curve! function, you will get data/fp.dat and data/ev.dat files, where they contain fixed points and eigenvalues, respectively.
function calc_fixed_point_vec(model_path::String)::Tuple{Array,Array}
global p = param_values()
new_curve!(
model_path, p, diffeq, get_derivatives, get_steady_state,
direction=false, bifparam=BP, n_state=SN
)
fp::Array = readdlm(joinpath(model_path, "data", "fp.dat"), '\t', Float64, '\n')
ev::Array = readdlm(joinpath(model_path, "data", "ev.dat"), '\t', Float64, '\n')
br::Array = get_bistable_regime(ev, SN)
return fp, br
endPlot results
function bifurcation_diagram(model_path::String, fp::Array, br::Array)
rc("figure", figsize=(8, 6))
rc("font", family="Arial")
rc("font", size=24)
rc("axes", linewidth=1)
rc("xtick.major", width=1)
rc("ytick.major", width=1)
rc("lines", linewidth=3)
plot(fp[1:br[1]-1, VN+1], fp[1:br[1]-1, V.E+1], "k-")
plot(fp[br, VN+1], fp[br, V.E+1], lw=1.5, "k--")
plot(fp[br[end]+1:end, VN+1], fp[br[end]+1:end, V.E+1], "k-")
xlabel("Serum (percentage)")
ylabel("E2F (μM)")
xlim(0, 2)
xticks([0, 0.5, 1, 1.5, 2])
yscale("log")
ylim(1e-4, 2)
yticks([1e-4, 1e-2, 1])
savefig(joinpath(model_path, "bifurcation_diagram.pdf"), bbox_inches="tight")
close()
endRun all functions defined above
const MODEL_PATH = "."
fp, br = calc_fixed_point_vec(MODEL_PATH);
bifurcation_diagram(MODEL_PATH, fp, br);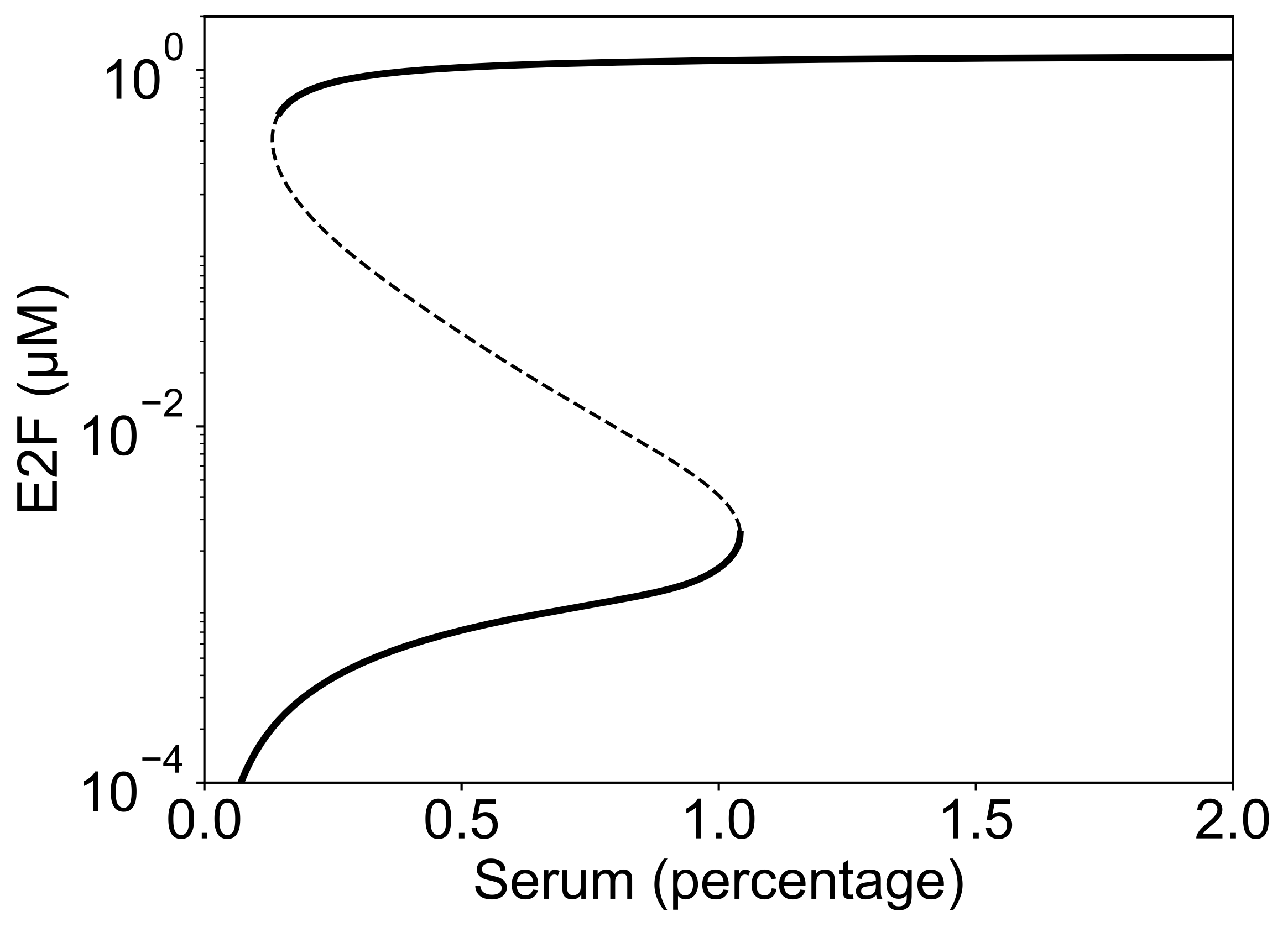
Stable (solid) and unstable (dashed) steady states of E2F activity with respect to serum stimulation.
For more examples, please refer to examples/bifurcation.