Circuitscape Documentation
Circuitscape is an open-source Julia program that uses circuit theory to model connectivity in heterogeneous landscapes. Its most common applications include modeling movement and gene flow of plants and animals, as well as identifying areas important for connectivity conservation. Circuit theory complements commonly-used connectivity models because of its connections to random walk theory and its ability to simultaneously evaluate contributions of multiple dispersal pathways. Landscapes are represented as conductive surfaces, with low resistances assigned to landscape features types that are most permeable to movement or best promote gene flow, and high resistances assigned to movement barriers. Effective resistances, current flow, and voltages calculated across the landscapes can then be related to ecological processes, such as individual movement and gene flow.
More detail about the underlying model, its parameterization, and potential applications in ecology, evolution, and conservation planning can be found in McRae (2006) and McRae et al. (2008).
More detail about implementation can be found in the Circuitscape In Julia paper.
Quick Start Guide
This is a quick start guide. If you're looking for basics on how to use Circuitscape, refer to the usage guide.
To run Circuitscape, you need to install the latest version of Julia. At the Julia prompt, install the Circuitscape package by running the following code
using Pkg
Pkg.add("Circuitscape")You can also install Circuitscape in a local project as well.
Running a job
A Circuitscape job is fully described by an INI file. This configuration file consists of file paths to data as well as flag values. A detailed list of fields found an in an INI File can be found in the Files and Flags section. Examples can be found in the test folder. You can also use a builtin UI to build Circuitscape jobs. For more on that, skip ahead to Building a Circuitscape Job.
If you do have your INI file handy, you can run the job by:
compute("myjob.ini")Building a Circuitscape Job

The builder is kicked off by calling the start() function from the Julia prompt. It will build an INI file for you step by step, and either run the job directly or write the final INI file to a specified location. You can exit the builder at any time by hitting Cntrl+C.
Please note that this version of Circuitscape does not come with a GUI.
You can, as a fallback, manually write your own INI file by copying and pasting an example from the test folder and change values as needed.
Citing Circuitscape
Please use the following to cite Circuitscape:
@article{Anantharaman2020,
doi = {10.21105/jcon.00058},
url = {https://doi.org/10.21105/jcon.00058},
year = {2020},
publisher = {The Open Journal},
volume = {1},
number = {1},
pages = {58},
author = {Ranjan Anantharaman and Kimberly Hall and Viral B. Shah and Alan Edelman},
title = {Circuitscape in Julia: High Performance Connectivity Modelling to Support Conservation Decisions},
journal = {Proceedings of the JuliaCon Conferences}
}What is new with Circuitscape?
The new Circuitscape is built entirely in the Julia language, a new programming language for technical computing. Julia is built from the ground up to be fast. As such, this offers a number of advantages over the previous version, and these are detailed below.
Faster and More Scalable
We benchmarked Circuitscape.jl (v0.1.0) with the Python version (v4.0.5) to obtain the following results. We started up Circuitscape with 16 parallel processes, and used benchmark problems from the standard Circuitscape benchmark suite.
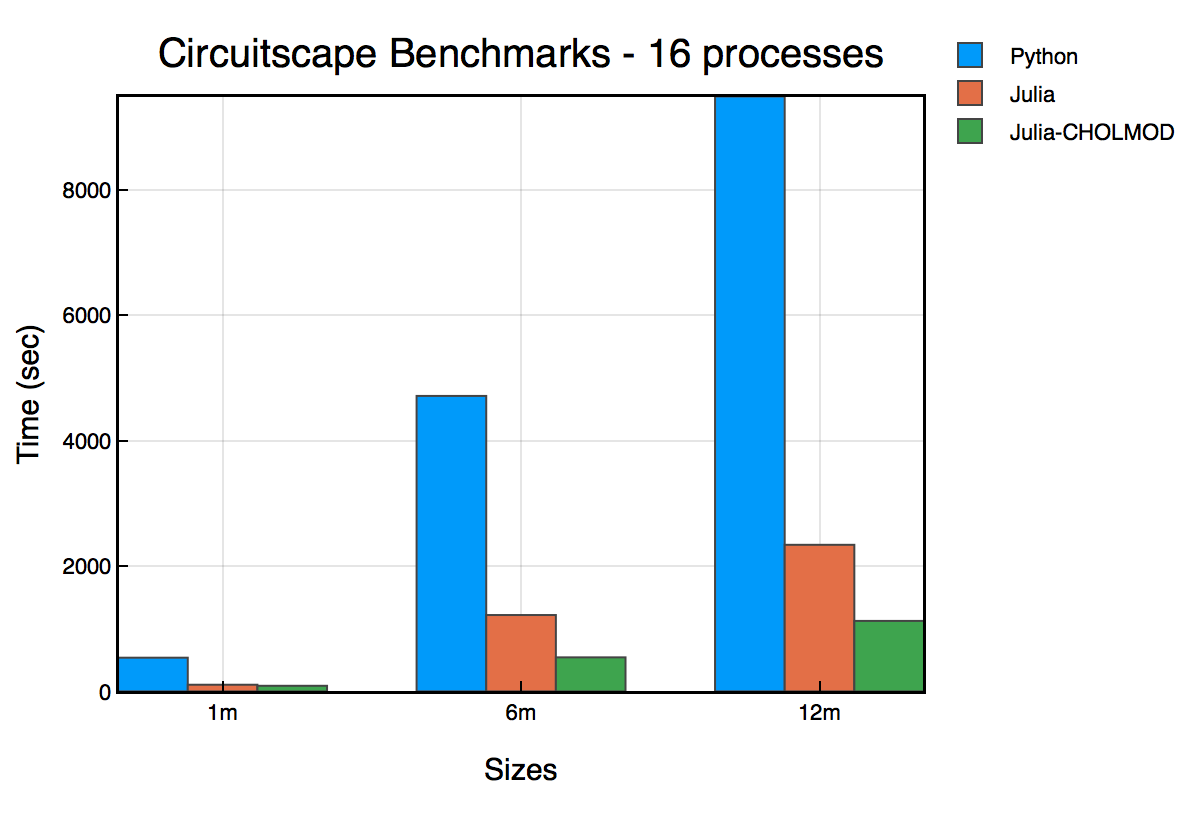
These benchmarks were run on a Linux (Ubuntu) server machine with the following specs:
- Name: Intel(R) Xeon(R) Silver 4114 CPU
- Clock Speed: 2.20GHz
- Number of cores: 20
- RAM: 384 GB
From the benchmark, we see that the new version is upto 4x faster on 16 processes. However, the best performing bar in the chart is Julia-CHOLMOD, which is a new feature introduced.
New Solver Mode - CHOLMOD
Julia-CHOLMOD is a new solver mode used in the new Circuitscape. It performs a cholesky decomposition on the graph constructed, and performs a batched back substitution to compute the voltages. It plugs into the CHOLMOD library, which is part of the SuiteSparse collection of high performance sparse matrix algorithms.
To use the this new mode, include a line in your Circuitscape INI file:
solver = cholmodThe cholesky decomposition is a direct solver method, unlike the algebraic multigrid method used by default in both the old and the new version. The advantage with this new direct method is that it can be much faster than the iterative solution, within a particular problem size.
Word of caution: The cholesky decomposition is not practical to use beyond a certain problem size because of phenomenon called fill-in, which results in loss of sparsity and large memory consumption.
Parallel, everywhere
The old Circuitscape had limited support for parallelism, which worked on Mac and Linux, but didn't work on Windows.
Julia as a programming language is built from the ground up to be parallel, and as a result the new Circuitscape natively supports parallelism on all three platforms.
Single Precision (Experimental)
The new Circuitscape introduces the ability to run problems in single precision as opposed to the standard double precision.
Single precision usually takes much less memory, but trades off against solution accuracy.
Use this new feature by including a line in your config file:
precision = singleRelated Projects
- Omniscape.jl - Omnidirectional connectivity analysis built on top of Circuitscape
- AlgebraicMultigrid.jl - Algebraic Multigrid methods in Julia. This is the default solver used in Circuitscape.
Further Reading
Beier, P., W. Spencer, R. Baldwin, and B.H. McRae. 2011. Best science practices for developing regional connectivity maps. Conservation Biology 25(5): 879-892
Dickson B.G., G.W. Roemer, B.H. McRae, and J.M. Rundall. 2013. Models of regional habitat quality and connectivity for pumas (Puma concolor) in the southwestern United States. PLoS ONE 8(12): e81898. doi:10.1371/journal.pone.0081898
McRae, B.H. 2006. Isolation by resistance. Evolution 60:1551-1561.
McRae, B.H. and P. Beier. 2007. Circuit theory predicts Gene flow in plant and animal populations. Proceedings of the National Academy of Sciences of the USA 104:19885-19890.
McRae, B.H., B.G. Dickson, T.H. Keitt, and V.B. Shah. 2008. Using circuit theory to model connectivity in ecology and conservation. Ecology 10: 2712-2724.
Shah, V.B. 2007. An Interactive System for Combinatorial Scientific Computing with an Emphasis on Programmer Productivity. PhD thesis, University of California, Santa Barbara.
Shah,V.B. and B.H. McRae. 2008. Circuitscape: a tool for landscape ecology. In: G. Varoquaux, T. Vaught, J. Millman (Eds.). Proceedings of the 7th Python in Science Conference (SciPy 2008), pp. 62-66.
Spear, S.F., N. Balkenhol, M.-J. Fortin, B.H. McRae and K. Scribner. 2010. Use of resistance surfaces for landscape genetic studies: Considerations of parameterization and analysis. Molecular Ecology 19(17): 3576-3591.
Zeller K.A., McGarigal K., and Whiteley A.R. 2012. Estimating landscape resistance to movement: a review. Landscape Ecology 27: 777-797.